Major advances in breast cancer treatment have significantly improved survival rates, but metastatic disease continues to be the leading cause of breast cancer-related deaths. Developing novel therapies for metastatic breast cancer requires preclinical breast cancer models that better-recapitulate important clinical features of the disease, including metastasis.
In this blog post we explore how to leverage Patient-Derived Xenograft (PDX) models for modeling metastatic breast cancer. We will look at models that spontaneously recapitulate the clinical metastatic process; a feature that is missing from conventional xenograft models.
Metastatic Breast Cancer
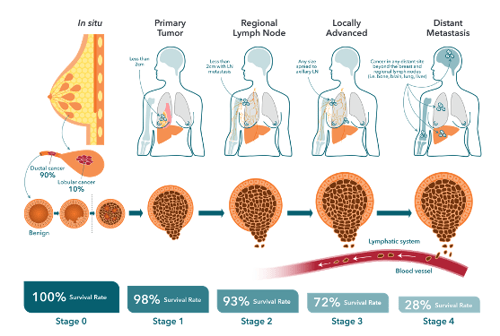
Breast cancer treatment typically strives to first reduce tumor burden followed by a course of adjuvant therapy aimed at preventing recurrences. Estimates indicate that up to 30% of early-stage breast cancers eventually progress to metastatic disease, where the primary tumor has metastasized beyond the breast to other parts of the body. This is usually in the form of a distant recurrence, sometimes occurring many years after original diagnosis and primary tumor resection.
The mechanisms leading to the production of disseminating tumor cells (DTC), their initial dormancy, and then later "reawakening" are poorly understood but have been shown to involve both genetic and epigenetic (genomic) changes in cancer cells, plus contributions from the tumor microenvironment (TME). The formation of a metastasis is a multistep process, requiring cancer cells to acquire specific properties to allow them to:
- Migrate from the primary tumor site,
- Survive in the lymphatic system or blood vessels,
- Colonize distant site(s) to form secondary tumors.
Clearly, there remains an unmet need for more effective and safe treatments for patients with metastatic breast cancer.
A major challenge in developing adjuvant breast cancer therapies has been a lack of patient-relevant in vivo preclinical models that adequately recapitulate key features of human tumors, which are slow growing and spontaneously metastatic.
Cell Line-derived Xenograft Models of Breast Cancer
Typically, researchers turn to conventional cell line-derived xenograft models for drug development studies. However, it is well-known that many drugs fail in the clinic despite highly promising preclinical evidence. While both cell line-derived syngeneic (i.e., developed in animals with a competent mouse immune system) and xenograft (i.e., developed in immunodeficient animals) models are commonly and successfully used to model the metastatic processes associated with various types of cancer, questions have been raised as to what extent cell lines recapitulate the biology of human tumors.
For instance, numerous studies have shown that some breast cancer cell lines are poorly metastatic in vivo, likely due to substantial changes that have occurred in these cells over many years of growing in 2D on plastic plates. To overcome this limitation, cell line metastasis models often use tail vein or intracardiac injections to induce tumor growth at different tissue sites. Such models sidestep the early stages of the metastatic cascade, and tend to model later stages of disease, as opposed to the spontaneous metastatic PDX models described below.
PDX Models of Metastatic Breast Cancer
 PDX models offer unique advantages in metastasis modeling. PDX models are developed by transplanting human breast tumor cells or tissue from either the primary tumor (collected during a lumpectomy or mastectomy) or from a metastatic lesion orthotopically into the mammary fat pads of immunocompromised mice, without ever culturing the cells.
PDX models offer unique advantages in metastasis modeling. PDX models are developed by transplanting human breast tumor cells or tissue from either the primary tumor (collected during a lumpectomy or mastectomy) or from a metastatic lesion orthotopically into the mammary fat pads of immunocompromised mice, without ever culturing the cells.
While the transplantation can also be done subcutaneously, which is less technically demanding and allows for efficient monitoring of tumor growth, experience has shown that orthotopic PDX models are better at mimicking metastasis as compared to subcutaneous models. Thus, orthotopic models are believed to more closely resemble the true tendency of human tumors to spontaneously metastasize. They also better recapitulate the entire underlying metastatic cascade, including locoregional extension, lymphatic and hematogenous metastasis, and the progression from primary to late-stage disease, including the ability to establish multi-site metastases.
Spontaneous metastases in orthotopic PDX breast cancer models often occur in human relevant sites such as the lung, lymph nodes, bone, ovaries, and the peritoneum. Furthermore, pairs of models can be generated from the same patient which cover primary and metastatic disease, as well as multiple models from a patient over the course of their progressive metastasis treatment (including disease sensitive/resistant models).
Methods have also been established for incorporating bioluminescent imaging markers in dissociated tumor cells from PDXs to allow for efficient in vivo and ex vivo detection of metastatic lesions from multiple secondary organs. Coupled with functional genomic screening, orthotopic PDX breast cancer models can also be used to identify genes that drive or suppress metastases.
Advantages and Challenges of PDX Breast Cancer Models
There are several advantages to using PDXs for modeling metastatic and non-metastatic breast cancer, including:
- Over the long term, PDXs highly recapitulate the histological, genomic, and architectural features of the parental tumor over successive passages, albeit some heterogeneity has been known to occur at P0 (the initial mouse passage).
- Since PDXs are derived directly from patients, all breast cancer subtypes are represented within PDX panels which are now commercially available. This includes the luminal A subtype, which are ER+ slow growing tumors. ER+ PDXs offer a distinct advantage in preclinical research since there is a shortage of estrogen-dependent or endocrine-resistant human breast cancer models.
- Serial transplantation allows any initially limited tissue supplies to be expanded.
- For drug development purposes, the growth of PDX models can be synchronized across a large cohort of animals, allowing for efficacy testing of drugs in vivo.
Researchers should also be aware of the challenges associated with the use of PDX models, including:
- High technical expertise is required for generating spontaneous PDX metastatic models.
- Establishing PDX models from scratch is time-consuming and researchers should take into consideration the time it will take to establish their desired model.
- To capture inter-patient heterogeneity observed in the clinic, researchers should consider using panels of PDX models, instead of single isolated models.
Conclusion
Metastatic breast cancer continues to be the leading cause of death in people with breast cancer. Novel preclinical models that better recapitulate the full spectrum of the metastatic process are more likely to produce translatable research that can improve clinical success. Therefore, careful consideration should be taken when selecting your metastatic breast cancer models. Leveraging PDX models for modeling metastatic breast cancer, provides a highly translational option for rapidly progressing metastatic disease drug discovery programs.
Please visit our website for more information on our PDX models. Watch this on-demand webinar "Preclinical Solutions for Drug Development Targeting Advanced Breast Cancer" by Dr. Rajendra Kumari for further insights.