![]() Learn how unique genetic events can be used to generate a “genetic fingerprint” to ID and QC murine tumor models.
Learn how unique genetic events can be used to generate a “genetic fingerprint” to ID and QC murine tumor models.
The Importance of Model Identification
The correct identification and quality control (QC) of research models and cell lines is vitally important in validating and replicating results. The huge impact of misidentified cell lines on both basic research and drug discovery programs is well-documented. It’s estimated that 18-36% of cell lines are misidentified or cross-contaminated across biomedical research.
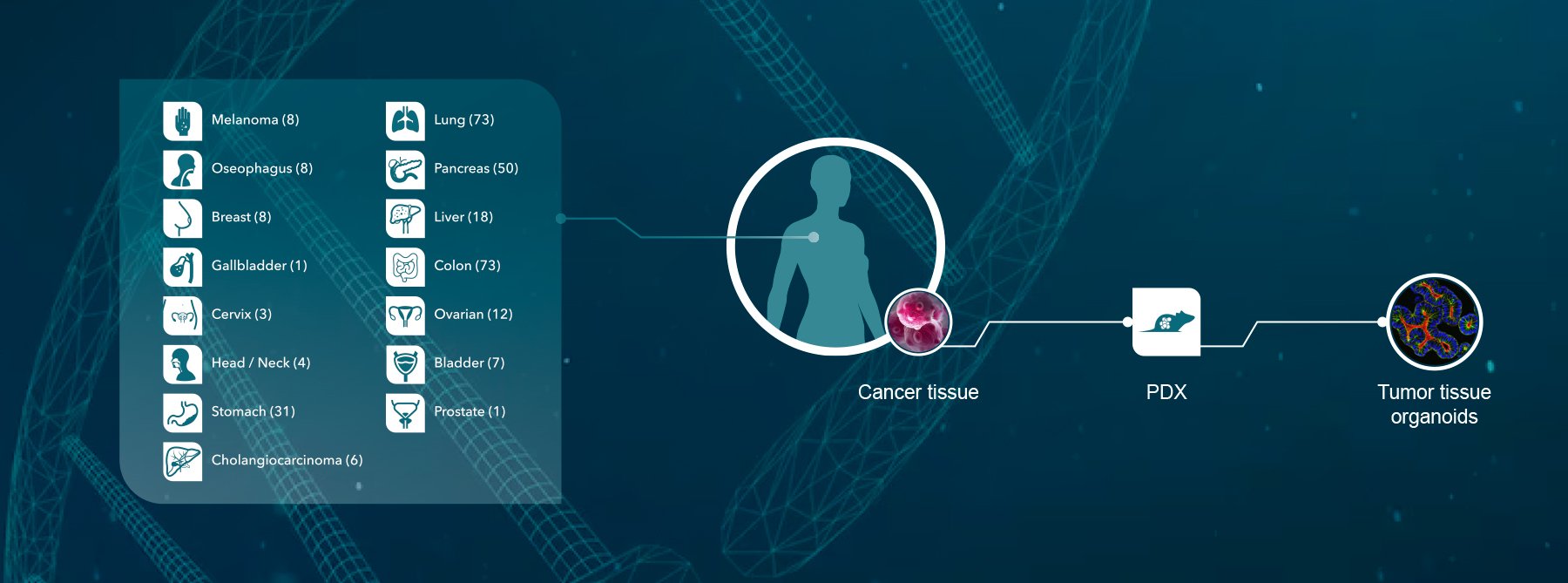
With human cell lines and xenograft models (such as conventional and patient-derived xenografts), tracking and genetic ID is reasonably straightforward. Short tandem repeat (STR) DNA profiling monitors core STR loci to:
- Show relatedness between cell lines.
- Uniquely identify human cells.
- Profile cell line comparisons.
Repositories such as the ATCC have compiled comprehensive databases of human cell line STR DNA profiles which can be used to match against in-house results.
Identification and Quality Control of Murine Tumor Models
For murine cell lines and newer tumor models such as syngeneics and tumor homografts, QC and ID is more tricky. Murine tumors originate from limited strains of inbred experimental mice. Therefore, they don’t have readily available markers to be used in ID assays like STR profiling or HLA typing.
In order to ID, track, and QC our murine models in house, we developed a novel and efficient assay where unique “genetic fingerprints” are established on an individual model basis.
Murine Tumor Model QC Process
The principle behind our QC process is to identify unique genetic features (e.g. unique gene fusions) within an individual tumor model. This is carried out using RNAseq on original cells and primary tumors.
The unique features are validated by RT-PCR and direct Sanger sequencing, and act as the model fingerprint moving forwards. Tumor homograft models developed from GEMM, or downstream passages of syngeneic cell lines/models are then analysed and validated against the parental fingerprint. This verifies that unique fusions are still present, and can monitor if new genetic events have occurred.
This allows our models continue to be correctly identified, and checks that the quality of the model is maintained.
Continuing Quality in Immuno-Oncology Models
Syngeneic models are being widely used across immunotherapy development, and new tumor homograft models continue to be generated. With the broad applications of these models across different labs in various studies, it’s prudent to track and QC the tumors by a routine assay to ensure quality and ID.
Using a “genetic fingerprint” developed using simple lab techniques provides an in-house assay that can be readily implemented to ensure model tracking from initial use or generation.