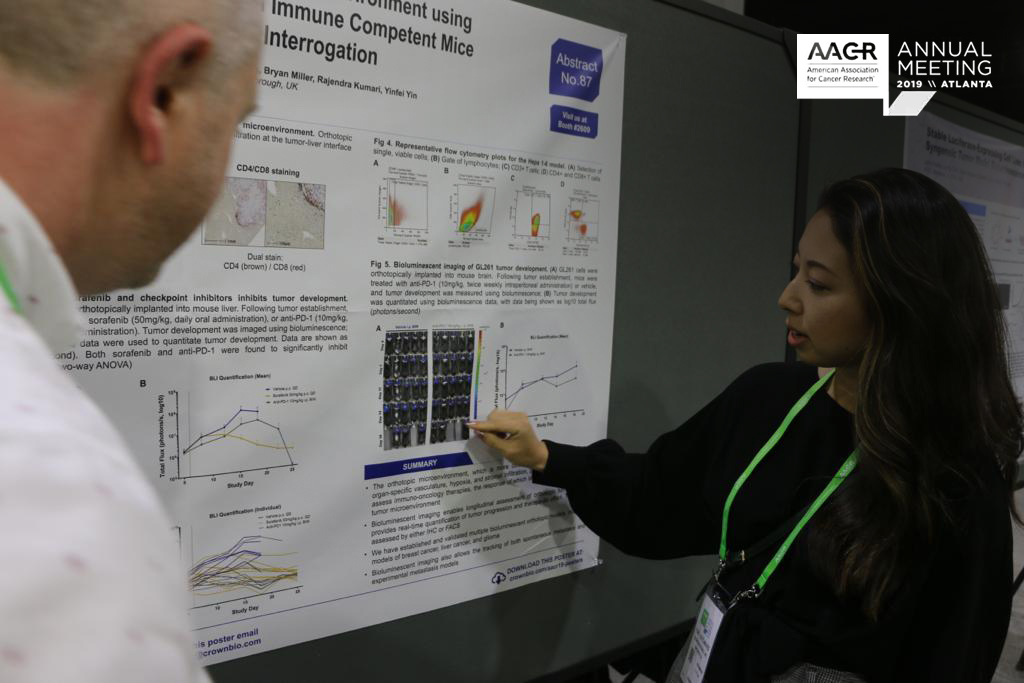
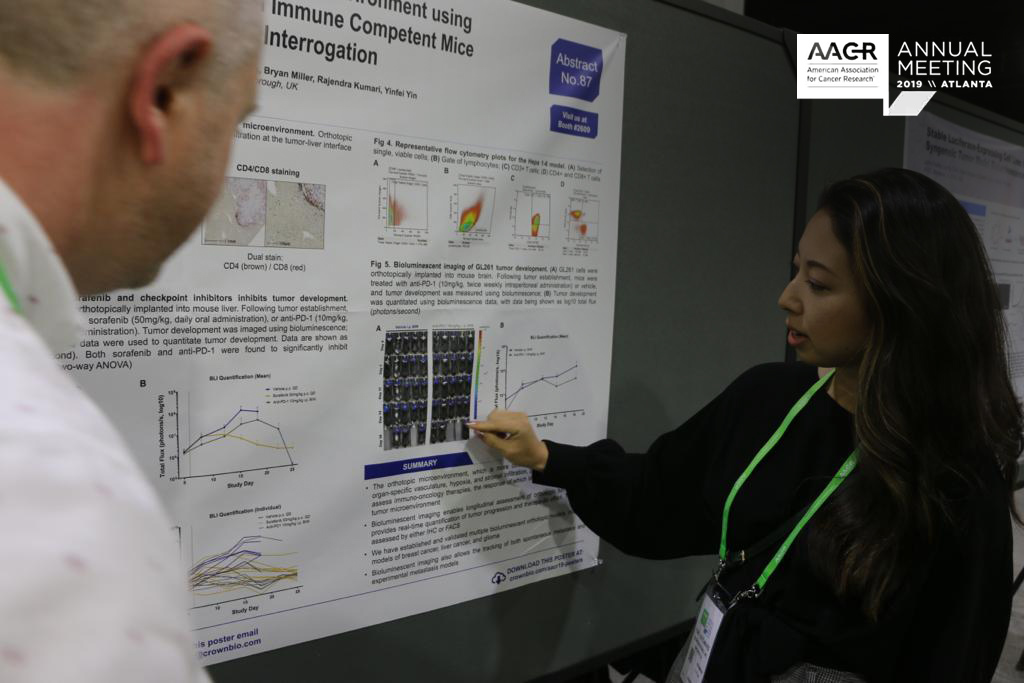
In Atlanta this week we’re presenting data on our new analysis platforms and preclinical technologies. This post recaps all the information on new CAR-T cell detection methods, an NGS-based SNP model identification array, data analysis platforms, and improvement on tumor growth mathematical model understanding.
In Atlanta this week we’re presenting data on our new analysis platforms and preclinical technologies. This post recaps all the information on new CAR-T cell detection methods, an NGS-based SNP model identification array, data analysis platforms, and improvement on tumor growth mathematical model understanding.
Sensitive Detection and Quantification of CAR-T Cells in Treated Patients (AACR 2019, Poster 1443)
 CAR-T cell therapy is a promising new personalized cancer treatment approach, using a patient’s own immune cells to fight cancer, and already FDA approved for treating certain types of NHL, DLBCL, and ALL.
CAR-T cell therapy is a promising new personalized cancer treatment approach, using a patient’s own immune cells to fight cancer, and already FDA approved for treating certain types of NHL, DLBCL, and ALL.
Despite being clinically used, there are still some questions to answer around CAR-T cell therapy. Notably, CAR-T cells need to be effectively monitored and characterized after administration to fully understand treatment effects and underlying mechanisms.
We’ve developed a new platform to detect and characterize rare CAR-T cells in patients. Our new platform looks to overcome the limitations of existing technologies - flow cytometry which has a low sensitivity, and real-time PCR which can’t be used to identify and characterize individual CAR-T cells.
CAR-T Cell Detection and Monitoring using RareCyte Technology
Our platform uses the RareCyte system which is validated for CAR-T cell detection in this poster. Preliminary data shows that our new method has a high sensitivity, at least 3 orders of magnitude above flow cytometry. A good linear range of quantification was seen, and single CAR-T cells could be picked for further single cell characterization.
Overall, this CAR-T cell detection and monitoring method could provide a great new opportunity for the clinic, in monitoring patient treatment, and understanding treatment mechanisms. There’s also the potential for diagnostic development, to provide guidance on patient treatment and prognosis.
A NGS-Based SNP Array for the Identification of Human Xenograft Tumors, Mouse Homograft Tumors, Human and Mouse Cell Lines, and Organoids (AACR 2019, Poster 3418)
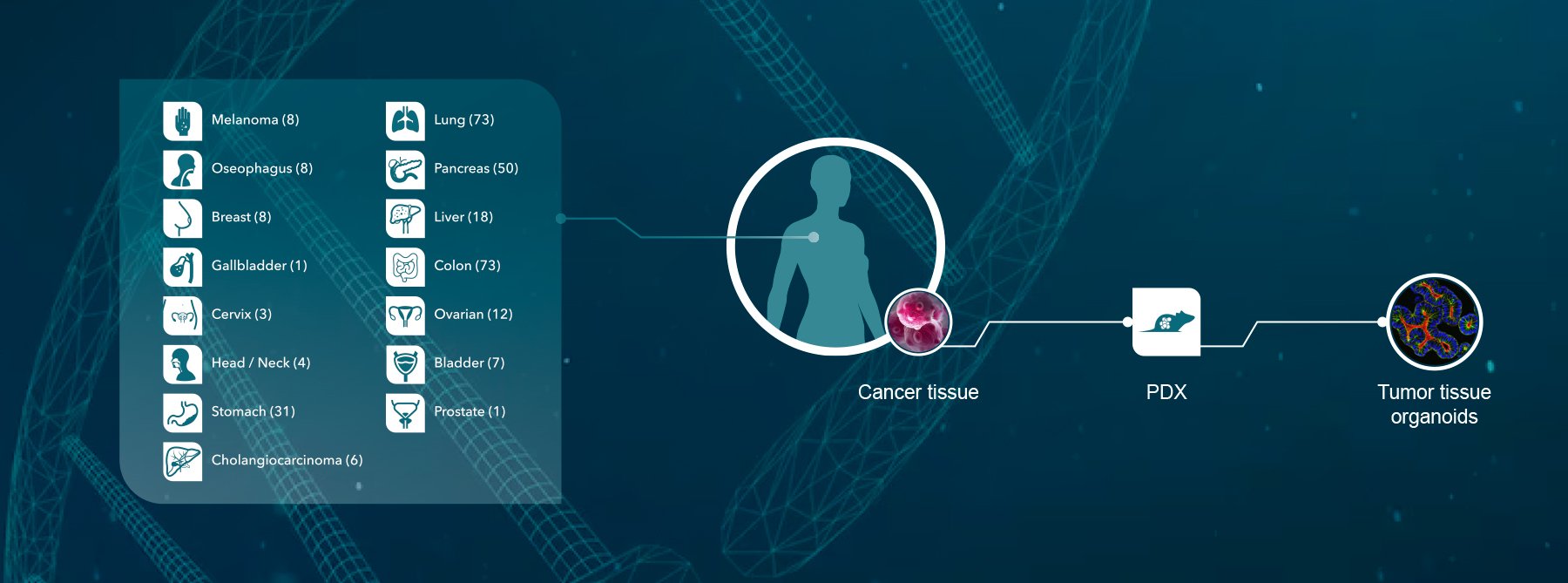
 To ensure authenticity of the variety of preclinical tumor models used in oncology drug development, models need to be tracked and quality controlled. Our next AACR poster presents a novel platform which can QC both human and murine models, including xenograft and patient-derived xenograft (PDX) tumors, mouse homograft tumors, human and mouse cell lines, and organoids.
To ensure authenticity of the variety of preclinical tumor models used in oncology drug development, models need to be tracked and quality controlled. Our next AACR poster presents a novel platform which can QC both human and murine models, including xenograft and patient-derived xenograft (PDX) tumors, mouse homograft tumors, human and mouse cell lines, and organoids.
The new technology is built on Single Nucleotide Polymorphism (SNP) based assays, which have been used in recent years to replace STR profiling of human cell lines and models. SNP-based assays have improved accuracy, can be automated, use low sample amounts and degraded samples, and multiple SNPs can be assayed simultaneously by Next Generation Sequencing (NGS) technology.
Multiple Functions of NGS-Based SNP Array
The poster details validation of our NGS-based SNP array for different functions including authenticating PDX tumor models, detecting viral infection and mycoplasma contamination, and characterizing common immunodeficient mouse strains. Unique fingerprints of around 200 SNPs are generated for each tumor model through RNAseq or WES data, and the poster also details our wide range of models for which fingerprints have been generated.
Overall, this platform provides an efficient and robust method for authenticating tumor models which can be adopted for the quality assurance of banking and tracking tumor model collections.
Mathematical Modeling of Tumor Growth in Mouse Models (AACR 2019, Poster 4613)
 Understanding the growth kinetics of tumor models is crucial for the accurate analysis of drug efficacy in preclinical cancer research. It’s known that different tumor models, like PDX, traditional xenografts, and syngeneics have different patterns of growth when being treated with vehicle or drugs.
Understanding the growth kinetics of tumor models is crucial for the accurate analysis of drug efficacy in preclinical cancer research. It’s known that different tumor models, like PDX, traditional xenografts, and syngeneics have different patterns of growth when being treated with vehicle or drugs.
Poster 4613 looked to answer the question of which mathematical models are best suited to describe tumor growth kinetics across different tumor models or for large numbers of tumors. While many models have been previously suggested for tumor growth kinetics, these two questions remain unanswered.
Evaluation of Eight Mathematical Models
We evaluated eight different mathematical models describing tumor growth kinetics, looking at vehicle and drug treatments using data from over 35,000 mice. The analysis covered conventional xenografts, PDX, and syngeneic models. Four goodness of fit criteria were used to assess the results.
Generally, we found that the mathematical models performed better for vehicle than drug groups, with mathematical model performance differing across the tumor models tested. The four goodness of fit criteria weren’t always consistent in assessing model performance, and overall we think further studies are needed to elucidate the applicability of the different mathematical models on individual datasets.
HuPharm: An Interactive Data Analysis Platform for Preclinical Studies (AACR 2019, Poster 5111)
 With preclinical data helping decide which new agents move to clinical development, it’s important that preclinical studies are carefully planned, properly executed, and correctly analysed for statistical relevance.
With preclinical data helping decide which new agents move to clinical development, it’s important that preclinical studies are carefully planned, properly executed, and correctly analysed for statistical relevance.
We’ve developed a new web platform to help with in vivo pharmacology data analysis and interpretation. Based upon user inputs, our HuPharm platform automatically generates appropriate figures and tables which facilitates preclinical data analysis. The intuitive and interactive interface is designed to help researchers (including those with less statistical training) on data validation, exploratory data analysis, and statistical analysis.
Poster 5111 details the packages used to build this platform, what an average analysis pipeline looks like, and the types of figures, tables, and analysis a user can be expect to receive in their end statistical report.
We hope this platform, with standardized statistical analysis reports, can help ensure consistency and to boost preclinical study productivity.
Read More on AACR 2019
For more on our AACR19 presentations, read our posts on PDX, humanized, and syngeneic models and research.