 CrownBio presented thirteen posters at AACR 2018, sharing our latest internal research into patient derived xenografts, humanized models, murine immuno-oncology models, and in vitro/ex vivo platforms. One particularly interesting poster we presented explored gut microbiota modulation following treatment with a PD-1 inhibitor, with potential implications for their use as therapeutic targets or biomarkers in immuno-oncology.
CrownBio presented thirteen posters at AACR 2018, sharing our latest internal research into patient derived xenografts, humanized models, murine immuno-oncology models, and in vitro/ex vivo platforms. One particularly interesting poster we presented explored gut microbiota modulation following treatment with a PD-1 inhibitor, with potential implications for their use as therapeutic targets or biomarkers in immuno-oncology.
AACR 2018 1264: Poster Commensal Microbiota is Involved in Anti-PD-1 mAb-Mediated Antitumor Activity in a Preclinical Tumor Model
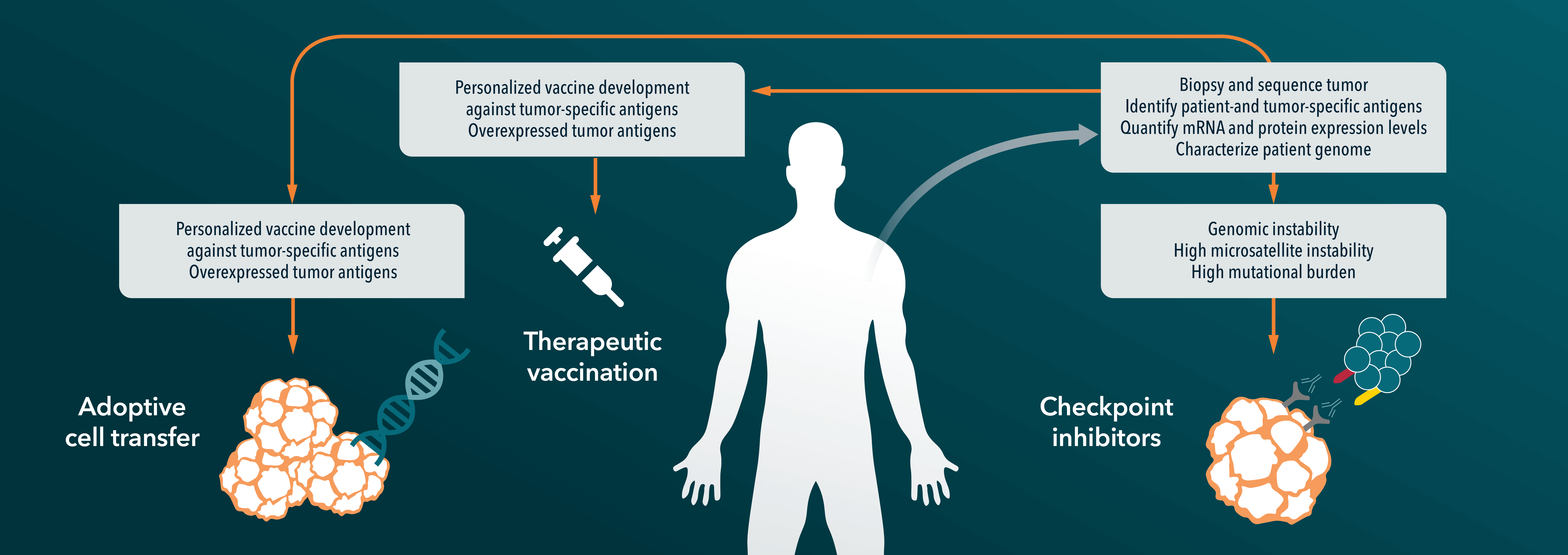
The gut microbiome is a hot topic in immuno-oncology, with a lot of active research into determining exactly how the microbiota affects immunotherapies via modulation of innate and adaptive immunity, and how to leverage this to improve response. Studies in preclinical mouse models have highlighted the key role of the gut microbiota in mediating tumor response to both chemotherapies and immune checkpoint inhibitors such as anti-PD-L1, PD-1, or CTLA-4.
 Multiple studies demonstrated that the gut microbiome can influence the outcome of PD-1 blockade in cancer patients or preclinical models, with several commensals shown to be associated with favorable clinical outcomes.
Multiple studies demonstrated that the gut microbiome can influence the outcome of PD-1 blockade in cancer patients or preclinical models, with several commensals shown to be associated with favorable clinical outcomes.
Expanding this research could lead to potential new therapeutic avenues. For our part, we studied the impact of anti-PD-1 treatment on gut microbiota makeup. We intended to determine whether the intestinal bacterial composition and abundance in tumor-bearing mice were associated with a specific immunotherapeutic outcome.
BALB/c mice bearing syngeneic CT-26 colon carcinomas were treated with five doses of an anti-PD-1 antibody, and a range of samples were collected pre- and post-treatment for 16S rRNA sequencing and data analysis.
Anti-PD-1 Treatment Influences the Composition of the Gut Microbiome
While tumor growth inhibition was observed for this model following anti-PD-1 treatment, a heatmap of relative abundance at genus level revealed a number of microbiota profile changes following treatment.
Looking specifically at the relative abundance of eight taxa at genus level, we can see statistically significant changes in abundance pre- and post-treatment for treated animals versus controls. LefSe analysis was also used to identify discriminative microbes at various levels.
Overall, this poster links gut microbiota modulation with potential immuno-microenvironment changes. If these microbiome changes can be consistently identified in treated animals, they have a potential to be used as biomarkers to predict response or as therapeutic targets in immuno-oncology.